RF Diffusion¶
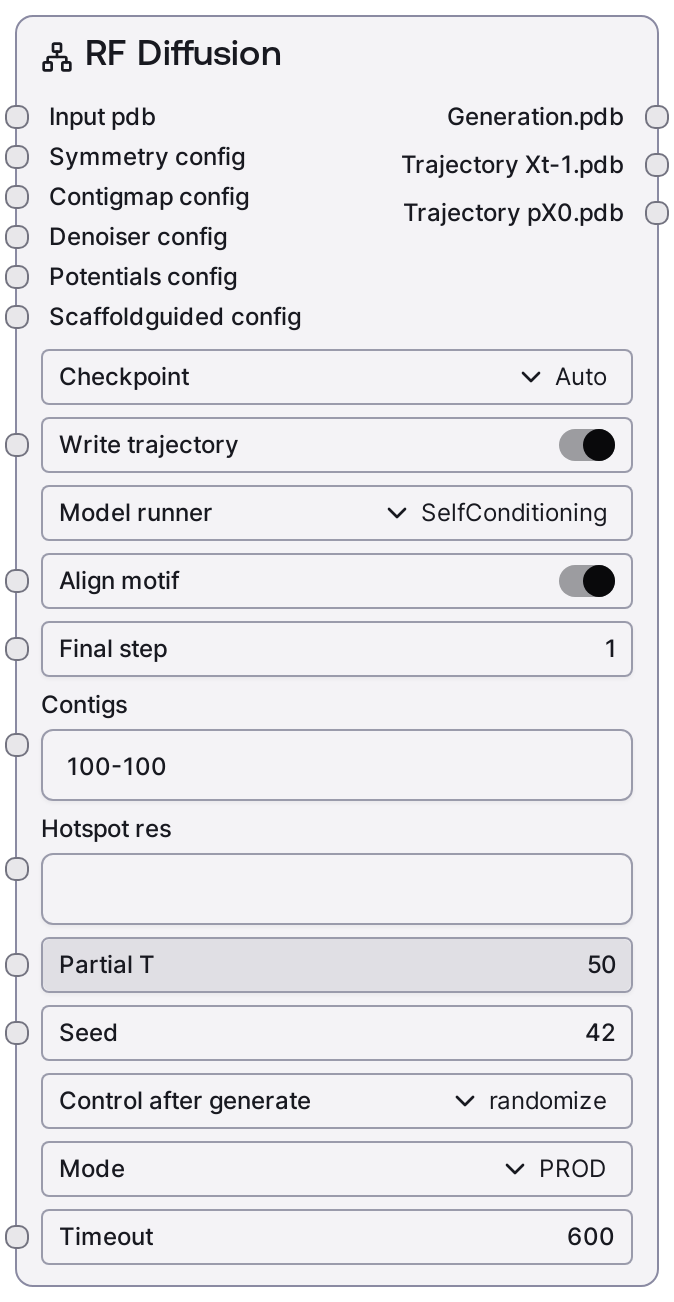
Usage¶
Use this node when you need to generate new protein backbones or redesign existing structures with RFdiffusion.
Typical scenarios: 1) Unconditional backbone generation by length - Leave input_pdb empty. - Set contigs to a purely numeric length or range (e.g. "120-120" or "80-100"). - Keep hotspot_res empty and avoid contigmap_config fields other than length.
2) Conditional design from an existing PDB - Provide input_pdb (monomer or complex). - Set contigs using fixed regions (e.g. "A10-25"), flexible ranges (e.g. "5-15"), and chain breaks ("/0") that match the PDB. - For partial diffusion (partial_T < 50), contigs must exactly describe the full residue layout of input_pdb.
3) PPI and binder design - Provide input_pdb containing target and binder chains. - Use contigs to define which regions are fixed vs flexible. - Optionally set hotspot_res as target residues and add potentials_config (from SaaSRFDiffusionPotentialsConfigNode) for interface-focused potentials.
4) Scaffold-guided / fold-conditioned design - Connect SaaSRFDiffusionScaffoldGuidedConfigNode to scaffoldguided_config with scaffold .pt files. - In scaffold-guided mode, contigs may be left empty and generated from scaffolds. - Optionally provide target_pdb and target .pt files via the same config for fold-conditioned binder design.
5) Advanced control via configuration nodes - Symmetry: SaaSRFDiffusionSymmetryConfigNode → symmetry_config for cyclic/dihedral/polyhedral symmetry. - Contigmap: SaaSRFDiffusionContigmapConfigNode → contigmap_config for inpainting and masking. - Denoiser: SaaSRFDiffusionDenoiserConfigNode → denoiser_config for noise schedules. - Potentials: SaaSRFDiffusionPotentialsConfigNode → potentials_config for energetic steering. - Scaffold-guided: SaaSRFDiffusionScaffoldGuidedConfigNode → scaffoldguided_config for topology constraints.
Pipeline position: - Upstream: PDB loader or prediction nodes; RF Diffusion Symmetry, Contigmap, Denoiser, Potentials, and Scaffold Guided Config nodes; scaffold *.pt loader nodes. - Downstream: structure viewers, scoring/evaluation, sequence extraction, docking, and simulation nodes.
Modes and recommendations: - Use mode="TEST" to debug quickly (reduced final_step, no trajectories, non-cautious) and mode="PROD" for full-quality runs. - Use mode="MOCK" when you need deterministic example outputs without calling the GPU service. - Disable write_trajectory unless you specifically need diffusion trajectories to save time and storage.
Inputs¶
| Field | Required | Type | Description | Example |
|---|---|---|---|---|
| checkpoint | True | CHOICE | RFdiffusion checkpoint to use. "Auto" lets the service pick a suitable model; other options (Base, Complex_base, Complex_Fold_base, InpaintSeq, InpaintSeq_Fold, ActiveSite, Base_epoch8, Complex_beta) force specific checkpoints. Ensure compatibility with your design task (e.g. complexes or inpainting). | Auto |
| write_trajectory | True | BOOLEAN | Whether to include diffusion trajectory PDBs (Xt-1 and pX0). Enabling this adds two outputs and increases runtime and data size. | True |
| model_runner | True | CHOICE | Sampler implementation used by RFdiffusion. Options: "default", "SelfConditioning" (common default), and "ScaffoldedSampler" (for scaffold-guided runs). | SelfConditioning |
| align_motif | True | BOOLEAN | Align the model’s predicted motif back to the input motif after diffusion. Recommended when preserving motif geometry in conditional design. | True |
| final_step | True | INT | Reverse diffusion step at which to stop (1–50). Larger values provide a fuller denoising schedule; smaller values speed up inference at potential cost to quality. In TEST mode this is internally set to 49. | 50 |
| contigs | True | STRING | Defines sequence layout for generation. In unconditional mode, use only numeric length ranges (e.g. "100-100"). In conditional mode, combine fixed motifs (e.g. "A10-25"), flexible ranges (e.g. "5-15"), and chain breaks ("/0"). For partial diffusion (partial_T != 50), contigs must exactly match the residue count and ordering of input_pdb. | 5-15/A10-25/30-40/0 B1-100 |
| partial_T | True | INT | Timestep to start denoising from (0–50). Set to 50 to disable partial diffusion (internally treated as no partial_T). Values <50 enable partial diffusion and require a valid input_pdb. | 50 |
| seed | True | INT | Base random seed (0–4294967295) used by RFdiffusion. The node also derives per-output seeds from this to make results reproducible. | 42 |
| mode | True | CHOICE | Execution mode. "PROD" runs the real RFdiffusion service with full settings. "TEST" reduces final_step, disables trajectory writing and cautious mode for faster debugging. "MOCK" returns predefined example results without calling the service. | PROD |
| timeout | True | INT | Timeout in seconds for the RFdiffusion process (30–12000). If the external service does not respond within this period, the node raises a timeout error. | 1200 |
| hotspot_res | False | STRING | Optional hotspot residues on the target protein, used for PPI and binder designs. Parsed as a list of residue identifiers and validated. Must be empty in unconditional sampling; in scaffold-guided mode, a target protein must be configured if hotspot_res is set. | A45,A48,A52 |
| input_pdb | False | PDB | Optional initial PDB structure used for conditional or partial diffusion. The node extracts the first PDB from this input. Required whenever partial_T < 50 or when contigs specify fixed motif regions that refer to an existing structure. | {'file_id': 'job-2024-10-01-structure-1', 'filename': 'input_complex.pdb', 'content': 'PDB_CONTENT_HANDLE'} |
| symmetry_config | False | RFDIFFUSION_SYMMETRY_CONFIG | Optional symmetry configuration from SaaSRFDiffusionSymmetryConfigNode. Controls symmetry type and options like recentering, radius, modeling neighbors, and symmetric self-conditioning for symmetric oligomer design. | {'recenter': True, 'radius': 12, 'model_only_neighbors': False, 'symmetric_self_cond': True, 'symmetry': 'c3'} |
| contigmap_config | False | RFDIFFUSION_CONTIGMAP_CONFIG | Optional contig-map configuration from SaaSRFDiffusionContigmapConfigNode. Defines sequence and structure inpainting (inpaint_seq, inpaint_str, inpaint_str_helix/strand/loop), regions to provide (provide_seq), and length. In unconditional mode, only length should be set; other fields must remain empty. | {'inpaint_seq': 'A30-50', 'inpaint_str': '', 'inpaint_str_helix': '', 'inpaint_str_strand': '', 'inpaint_str_loop': '', 'provide_seq': '1-29,51-100', 'length': '100-110'} |
| denoiser_config | False | RFDIFFUSION_DENOISER_CONFIG | Optional denoiser configuration from SaaSRFDiffusionDenoiserConfigNode. Adjusts noise scales for C-alpha translations and frame rotations at initial and final steps, with linear or constant schedules, to control exploration vs stability. | {'noise_scale_ca': 1.0, 'final_noise_scale_ca': 0.5, 'ca_noise_schedule_type': 'linear', 'noise_scale_frame': 1.0, 'final_noise_scale_frame': 0.5, 'frame_noise_schedule_type': 'linear'} |
| potentials_config | False | RFDIFFUSION_POTENTIALS_CONFIG | Optional guiding potentials configuration from SaaSRFDiffusionPotentialsConfigNode. Specifies guiding_potentials (e.g. monomer_ROG, interface_ncontacts, olig_contacts, substrate_contacts) plus global guide_scale, guide_decay, oligomer contact flags, and optional ligand substrate name. | {'guiding_potentials': ['type:monomer_ROG,weight:1,min_dist:15', 'type:interface_ncontacts,weight:1,r_0:8,d_0:6'], 'guide_scale': 10, 'guide_decay': 'linear', 'olig_inter_all': False, 'olig_intra_all': False, 'olig_custom_contact': 'A!B,B&C', 'substrate': 'ATP'} |
| scaffoldguided_config | False | RFDIFFUSION_SCAFFOLDGUIDED_CONFIG | Optional scaffold-guided/fold conditioning configuration from SaaSRFDiffusionScaffoldGuidedConfigNode. Enables topology-constrained design using scaffold secondary-structure/adjacency *.pt files, and can include a target protein and its *.pt files for fold-conditioned PPI design. When scaffoldguided is true, contigs may be omitted and auto-generated from scaffold structure. | {'scaffoldguided': True, 'scaffold_secstruc_adj': {'type': 'cloud_path', 'provider': 'gcs', 'cloud_folder_path': 'gs://my-bucket/rfdiffusion/scaffolds/'}, 'scaffold_list': ['scaffold_1', 'scaffold_5'], 'sampled_insertion': 5, 'sampled_N': 3, 'sampled_C': 3, 'ss_mask': 2, 'systematic': False, 'mask_loops': True} |
Outputs¶
| Field | Type | Description | Example |
|---|---|---|---|
| generation.pdb | PDB | Final designed or generated protein backbone from RFdiffusion. This is the primary result used for downstream visualization, scoring, sequence extraction, or further modeling. It includes metadata such as seed and a folder_name of "rf_diffusion" for organization. | {'file_id': 'rfdesign-2024-10-01-001', 'filename': 'rf_diffusion_design.pdb', 'content': 'PDB_CONTENT_HANDLE', 'metadata': {'seed': 42, 'folder_name': 'rf_diffusion'}} |
| trajectory_Xt-1.pdb | PDB | Optional trajectory PDB holding Xt-1 structures along the diffusion path. Returned only when write_trajectory is true. Useful for analyzing the denoising trajectory or diagnosing convergence issues. | {'file_id': 'rfdesign-2024-10-01-001-traj-Xt-1', 'filename': 'rf_trajectory_Xt-1.pdb', 'content': 'PDB_TRAJECTORY_CONTENT', 'metadata': {'seed': 43, 'folder_name': 'rf_diffusion'}} |
| trajectory_pX0.pdb | PDB | Optional trajectory PDB containing pX0 predictions over diffusion steps, also only present when write_trajectory is true. Use this to inspect the evolution of predicted denoised structures across the schedule. | {'file_id': 'rfdesign-2024-10-01-001-traj-pX0', 'filename': 'rf_trajectory_pX0.pdb', 'content': 'PDB_TRAJECTORY_CONTENT', 'metadata': {'seed': 44, 'folder_name': 'rf_diffusion'}} |
Important Notes¶
- Performance: RFdiffusion runs are computationally expensive, especially for long sequences, complexes, and heavy constraints. Use TEST mode and disable trajectories while iterating, then switch to PROD with trajectories only when needed.
- Limitations: Unconditional mode requires contigs with only numeric ranges and forbids hotspot_res and most contigmap_config fields. Conditional and partial diffusion modes require contigs consistent with the input_pdb chain and residue layout, otherwise explicit validation errors are raised.
- Behavior: A partial_T value of 50 is treated as no partial diffusion (internally converted to None). For partial_T < 50, input_pdb is mandatory; if missing, the node raises an error before calling the service.
- Behavior: When scaffoldguided_config enables scaffoldguided mode, contigs may be None or empty and are auto-generated from scaffolds. In this case, hotspot_res is only valid if a target protein is configured in the same scaffoldguided_config.
- Behavior: In TEST mode, the node modifies inference parameters (final_step forced to 49, write_trajectory set to false, cautious mode disabled), so results will not exactly match PROD even with identical seeds.
- Limitations: Structured string inputs (contigs, guiding_potentials, olig_custom_contact, hotspot_res) must follow the documented formats; the node validates them and fails fast rather than attempting to silently fix malformed values.
Troubleshooting¶
- Unconditional contigs error: If you see an error like "For unconditional sampling, expected
contigsto specify only length range ...", you are using chain letters or motifs without an input_pdb. Fix by using numeric-only ranges (e.g. "120-120") and clearing hotspot_res and contigmap_config fields (except length). - Partial diffusion without input_pdb: Error "For partial diffusion sampling, expected
input_pdbto be provided." occurs when partial_T < 50 but no input_pdb is given. Either set partial_T to 50 to disable partial diffusion or supply a valid input_pdb and matching contigs. - Conditional contigs missing fixed regions: An error like "For conditional sampling, expected contigs to specify least one fixed region ..." means you passed input_pdb but contigs only contains numeric gaps. Add at least one fixed motif region (e.g. "A10-25") or remove input_pdb to switch to unconditional mode.
- Hotspot residues without target: If hotspot_res is specified while scaffoldguided_config has no target protein configured, the node raises an error explaining that hotspots require a target. Fix by adding a target_pdb in SaaSRFDiffusionScaffoldGuidedConfigNode or clearing hotspot_res.
- Timeout from RFdiffusion service: When runs fail due to timeout, increase the timeout parameter, simplify contigs or constraints, disable trajectories, or test with smaller systems first. Use TEST mode to verify connectivity and configuration before long PROD runs.