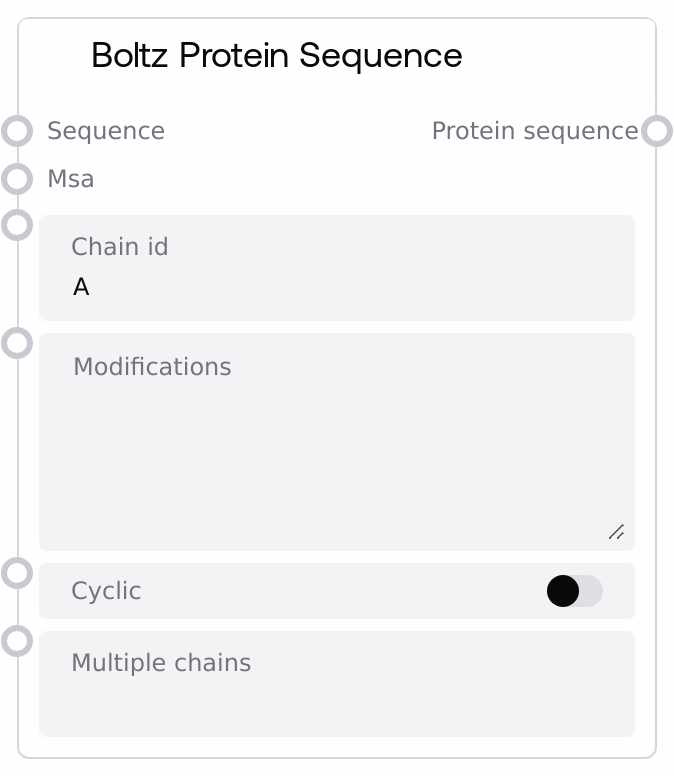
Boltz Protein Sequence
This node converts a protein FASTA input into a structured protein sequence specification for Boltz YAML workflows. It parses the FASTA to extract the amino acid sequence and an optional header-based sequence name, attaches optional MSA data and residue-level modifications, and can mark the protein as cyclic. It also supports assigning the same sequence to multiple chain IDs, returning a list-wrapped protein entry ready to be combined into a full Boltz configuration.
Usage
Use this node when you need to define one or more protein chains for a Boltz-based modeling run, such as structure prediction, protein–ligand complexes, or multimers. Provide a protein sequence in FASTA format, optionally connect an MSA, specify any residue modifications, and indicate if the protein is cyclic. The chain_id sets the primary chain, while multiple_chains lets you duplicate the same protein on additional chains (e.g., for homodimers). In a typical workflow, you (1) feed your FASTA into Boltz Protein Sequence, (2) optionally connect an MSA source, (3) configure modifications and cyclic if needed, (4) send the protein_sequence output into a Boltz List Combiner along with other sequence/constraint/template/property objects, (5) send the combined list into a Boltz YAML Combiner, and (6) pass the resulting boltz_yaml and boltz_files into Boltz Predict. Pair this node with Boltz Ligand Sequence for ligands, Boltz Constraint Builder for restraints, Boltz Template Builder for structural templates, and Boltz Property Builder for affinity or other property predictions.
| Field | Required | Type | Description | Example |
|---|
| chain_id | True | STRING | Primary chain identifier for this protein in the Boltz configuration. Must be a non-empty string and should be unique across all sequences in the final YAML (for example A, B, L). If this ID clashes with any other chain ID, the Boltz YAML Combiner will later raise a duplicate-chain error. | A |
| sequence | True | FASTA | Protein sequence in FASTA format. The value is treated as FASTA-like text: if it starts with '>', the first header line is used to derive a sequence name and subsequent lines are concatenated as the amino acid sequence. If there is no header line, the content is treated as a raw sequence and a default name is used. The final sequence must be non-empty, otherwise the node will error. | >SARS_CoV2_Spike_RBD RVQPTESIVRFPNITNLCPFGEVFNATRFASVYAWNRKRISNCVADYSVLYNSASFSTFKCY |
| msa | False | MSA | Optional multiple sequence alignment for this protein, typically in A3M or CSV-like form. When provided, the node infers the content and format and attaches both to the protein object. If omitted or empty, downstream nodes will treat it as an "empty" MSA. For best results, the MSA should correspond to the same protein sequence. | >seq1 RVQPTESIVRFPNITNLCPFGEVFNATRFASVY >seq2 RVQPTESIVRFPDITNLCPFGEIFNATRFASVY |
| modifications | False | STRING | Residue-level modifications, one per line, using the format "position:ccd_code". Position is a 1-based integer index into the protein sequence; ccd_code is a three-letter code or similar identifier understood by the backend. Lines that do not contain a colon or that use a non-integer position are skipped with a warning; valid lines are turned into a list of {"position": int, "ccd": string}. | 5:MSE 42:SEP |
| cyclic | False | BOOLEAN | If true, marks the protein as cyclic in the Boltz specification, indicating that the N- and C-termini should be treated as connected. If false, the protein is treated as linear. | true |
| multiple_chains | False | STRING | Comma-separated list of additional chain IDs that should be identical copies of this protein sequence (for example B,C for a homodimer). The node combines chain_id and these IDs into a single id field that is either a list of chain IDs or a single string. All IDs must be unique across the entire Boltz YAML; duplicates will cause a validation error in the Boltz YAML Combiner. | B,C |
Outputs
| Field | Type | Description | Example |
|---|
| protein_sequence | * | A list containing a single protein sequence specification object in Boltz format. Its structure is [ { "protein": { "id": "A" or ["A", "B", ...], "sequence": "", "_sequence_name": "", "msa": , "_msa_format": "a3m" or "csv", and optionally "modifications": [{"position": int, "ccd": "code"}], "cyclic": true/false } } ]. This list is intended to be merged with other Boltz objects using Boltz List Combiner and then passed into Boltz YAML Combiner. | [{"protein": {"id": ["A", "B"], "sequence": "RVQPTESIVRFPNITNLCPFGEVFNATRFASVY", "_sequence_name": "SARS_CoV2_Spike_RBD", "msa": "msa_1.a3m", "_msa_format": "a3m", "modifications": [{"position": 5, "ccd": "MSE"}], "cyclic": false}}] |
Important Notes
- Performance: Supplying very large MSAs can significantly increase runtime and memory consumption in downstream Boltz Predict steps; consider trimming or subsampling alignments if performance is a concern.
- Limitations: The input sequence must resolve to a non-empty amino acid string. A header-only FASTA or whitespace-only input will cause validation errors such as "Protein sequence is required" or "Protein sequence cannot be empty".
- Behavior: When multiple_chains is set, the id field of the protein becomes a list of chain IDs. Later, Boltz YAML Combiner enforces that every chain ID across all sequences (proteins, nucleic acids, ligands) is globally unique and will fail on any duplicate.
- Behavior: Modification lines that do not match the "position:ccd_code" pattern, or whose position cannot be parsed as an integer, are ignored with a warning rather than stopping the workflow. Always double-check modification syntax if expected modifications are missing from outputs.
Troubleshooting
- Common Error 1 – 'Protein sequence is required': This occurs when the sequence input is missing or contains only whitespace. Ensure that the FASTA text includes actual sequence lines below the header, or that the raw sequence is provided without an empty header.
- Common Error 2 – 'Protein sequence cannot be empty': The node detected a FASTA header (starting with '>') but no sequence lines after it. Add one or more lines of amino acid sequence under the header.
- Common Error 3 – Duplicate chain ID errors in downstream nodes: If Boltz YAML Combiner reports a duplicate chain ID, another sequence node is using the same chain_id or one of the IDs in multiple_chains. Adjust IDs so that each chain across all sequences is unique.
- Common Issue – Missing or partial modifications: If some modifications do not appear in the output, check each line for the exact "position:ccd_code" format with a numeric position. Incorrect lines are silently skipped except for log warnings.