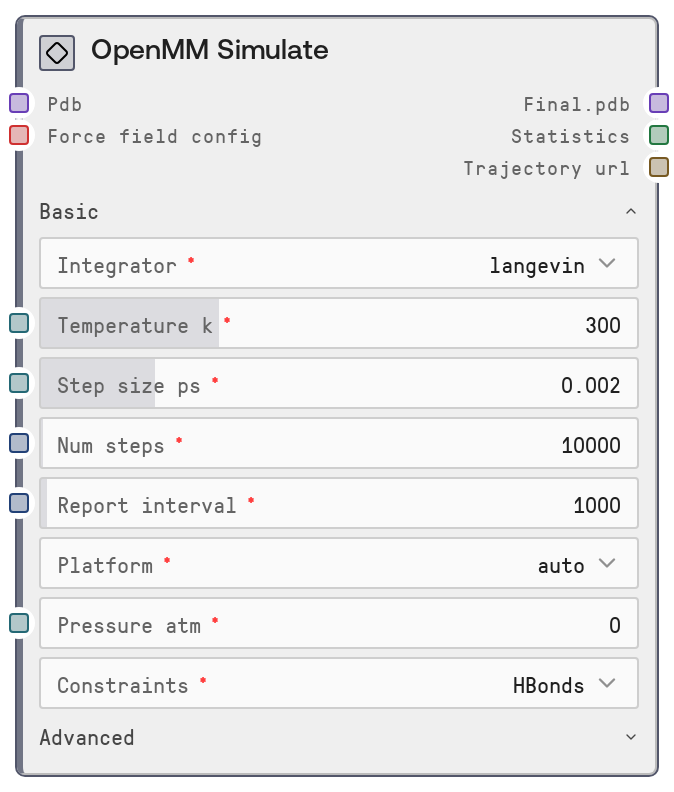
OpenMM Simulate
OpenMM Simulate performs molecular dynamics (MD) simulations via the salt-openmm-service for one or more input PDB structures. It uses a shared OpenMM force field configuration, supports multiple integrators and ensemble settings, and exposes controls for timestep, pressure, constraints, and nonbonded parameters. The node outputs final-frame PDBs, a combined statistics table from OpenMM’s StateDataReporter, and URLs to DCD trajectory files for downstream analysis.
Usage
Use this node when you need to run equilibrium or production MD simulations on biomolecular systems prepared earlier in your Salt workflow. Typical workflow: structure prediction/import → OpenMM ForceField Config → OpenMM Solvate → OpenMM Energy Minimize → OpenMM Simulate. Upstream nodes usually provide solvated, minimized PDBs and a force_field_config; downstream nodes can consume the final PDB structures, the statistics DataFrame, or fetch trajectories from the returned URLs for analysis (e.g., RMSD, clustering, secondary structure). For production runs, use integrator="langevin", temperature_k≈300, pressure_atm=1.0, appropriate constraints, and a report_interval such that num_steps / report_interval ≤ 50000 to satisfy the node’s row limit. Prefer platform="auto" with platform_precision="mixed" where GPUs are available, and start with conservative settings (step_size_ps≈0.002, HBonds constraints, standard PME tolerances) to ensure stability before optimizing performance.
| Field | Required | Type | Description | Example |
|---|
| pdb | True | PDB | Dictionary mapping PDB identifiers to PDB-formatted text for the structures to simulate. Must contain at least one entry; each value is a complete PDB string. Structures should normally be solvated and energy-minimized upstream for stable MD. | {'1-proteinA': 'ATOM 1 N MET A 1 11.104 13.207 7.235 1.00 20.00 N\nATOM 2 CA MET A 1 12.560 13.400 7.500 1.00 20.00 C\nTER\nEND\n'} |
| force_field_config | True | OPENMM_FORCE_FIELD_CONFIG | Force field and water model configuration produced by the OpenMM ForceField Config node. Must be a compatible combination (e.g., amber19-all.xml with amber19/tip3pfb.xml) as validated there. | {'force_field': 'amber19-all.xml', 'water_model': 'amber19/tip3pfb.xml'} |
| integrator | True | STRING (enum: langevin\|verlet\|brownian) | Integrator algorithm. "langevin" runs NVT with a thermostat (recommended for equilibrium sampling), "verlet" runs NVE energy-conserving dynamics, and "brownian" uses overdamped Langevin dynamics. | langevin |
| temperature_k | True | FLOAT | Simulation temperature in Kelvin (0.0–1000.0). 300.0 K is typical physiological temperature; 310.0 K approximates 37 °C. Used by thermostatted integrators and barostats. | 300.0 |
| step_size_ps | True | FLOAT | Integrator timestep in picoseconds (0.0001–0.01). A value of 0.002 ps (2 fs) is standard with HBonds constraints; larger steps may improve performance but can cause instability if too large for the chosen constraints and hydrogen masses. | 0.002 |
| num_steps | True | INT | Total number of MD integration steps (1–10000000). The simulated time is num_steps × step_size_ps; for example, 500000 steps at 0.002 ps equals 1000 ps (1 ns). | 500000 |
| report_interval | True | INT | Interval in steps at which trajectory frames and energy statistics are written (1–100000). The number of statistics rows per trajectory is approximately num_steps / report_interval and must be ≤ 50000 or the node will error. | 1000 |
| platform | True | STRING (enum: auto\|CUDA\|OpenCL\|CPU) | Compute platform for OpenMM. "auto" selects the fastest available platform. "CUDA" and "OpenCL" use compatible GPUs, while "CPU" forces CPU-only execution, which is slower but more universally available. | auto |
| pressure_atm | True | FLOAT | Target pressure in atmospheres (0.0–10.0). 0.0 runs NVT (constant volume), while 1.0 atm enables NPT (constant pressure) with box volume adjustments, recommended for solvated production simulations. | 1.0 |
| constraints | True | STRING (enum: HBonds\|AllBonds\|HAngles\|None) | Which internal coordinates to constrain. "HBonds" (default) constrains bonds to hydrogens and supports 2 fs timesteps. "AllBonds" with hydrogen mass repartitioning allows 4–5 fs timesteps. "HAngles" constrains angles involving hydrogens. "None" uses no constraints (slower and less stable). | HBonds |
| friction_coeff | False | FLOAT | Friction (collision) frequency for Langevin and Brownian integrators in 1/ps (0.01–100.0). Controls coupling to the heat bath; 1.0 ps⁻¹ is a common value for biomolecular simulations. | 1.0 |
| barostat_interval | False | INT | Number of steps between barostat volume moves (1–1000). Only used when pressure_atm > 0. A value of 25 is standard for NPT simulations. | 25 |
| nonbonded_cutoff_nm | False | FLOAT | Cutoff distance for short-range non-bonded interactions in nanometers (0.5–1.4). AMBER simulations often use 1.0 nm; CHARMM simulations often use 1.2 nm. The cutoff must be less than half the simulation box edge length. | 1.0 |
| hydrogen_mass | False | FLOAT | Hydrogen mass in atomic mass units for mass repartitioning (0.0–4.0). 0.0 disables repartitioning. Values around 3–4 amu, used with AllBonds constraints, allow larger timesteps (4–5 fs) for higher throughput; test stability before production use. | 3.0 |
| platform_precision | False | STRING (enum: auto\|single\|mixed\|double) | Precision mode for GPU platforms. "mixed" is recommended (single-precision forces, double-precision integration). "single" is fastest but less stable; "double" is most accurate but slowest. "auto" lets the backend choose a sensible default. | mixed |
| ewald_error_tolerance | False | FLOAT | Error tolerance for PME electrostatics (0.0001–0.001). Smaller values yield more accurate long-range electrostatics but reduce performance. 0.0005 is a common default. | 0.0005 |
| constraint_tolerance | False | FLOAT | Tolerance for the constraint solver (e.g., SHAKE/SETTLE) (1e-6–1e-4). Lower values enforce more accurate constraints at some performance cost. A typical choice is 1e-5. | 1e-05 |
Outputs
| Field | Type | Description | Example |
|---|
| final.pdb | PDB | Dictionary mapping each input pdb_id to the final simulation snapshot in PDB format. Each value is a PDB-formatted string representing the last frame of that trajectory, suitable for inspection or further refinement and analysis. | {'1-proteinA': 'ATOM 1 N MET A 1 11.352 13.489 7.102 1.00 20.00 N\nATOM 5234 O HOH W 700 32.110 25.430 19.876 1.00 20.00 O\nTER\nEND\n'} |
| statistics | DATAFRAME | Combined statistics table built from OpenMM StateDataReporter CSV outputs for all simulated structures. Includes a leading pdb_id column, all standard reporter columns (e.g., step, time, potentialEnergy, totalEnergy, temperature, volume, density), plus a trajectory_url column indicating the trajectory file associated with each row. | pdb_id,step,time(ps),potentialEnergy(kJ/mole),totalEnergy(kJ/mole),temperature(K),volume(nm^3),density(g/mL),trajectory_url 1-proteinA,0,0.0,-25743.2,-25510.8,298.7,125.6,1.02,https://s3-region.amazonaws.com/salt-md-trajectories/proteinA_run1.dcd 1-proteinA,1000,2.0,-25680.5,-25450.3,300.1,125.8,1.02,https://s3-region.amazonaws.com/salt-md-trajectories/proteinA_run1.dcd |
| trajectory_url | STRING | Plaintext string containing one trajectory URL per line, in the same order as the input pdb keys. Each URL typically points to a DCD file stored in object storage. For a structured pdb_id-to-URL mapping, use the trajectory_url column in the statistics output instead of parsing this string. | https://s3-region.amazonaws.com/salt-md-trajectories/proteinA_run1.dcd https://s3-region.amazonaws.com/salt-md-trajectories/proteinB_run1.dcd |
Important Notes
- Performance: The node sets a backend timeout of up to 5400 seconds (90 minutes) per call. Very long simulations or huge systems may approach this limit; consider splitting runs into shorter segments or reducing system size or num_steps.
- Performance: A hard limit of 50000 statistics rows per node call is enforced. The node checks num_steps / report_interval and raises an error if it exceeds 50000; increase report_interval or reduce num_steps to stay within this bound.
- Limitations: The pdb input must be a non-empty dictionary; an empty dict triggers a validation error. Additionally, poorly prepared structures (not solvated, not minimized, or with severe clashes) may cause the backend OpenMM simulation to fail or produce NaNs even if the node itself accepts the inputs.
- Behavior: When hydrogen_mass is 0.0, hydrogen mass repartitioning is effectively disabled and not sent to the backend. Use non-zero hydrogen_mass only with compatible constraints (typically AllBonds) and carefully validated timesteps.
- Behavior: The trajectory_url output is a convenience plaintext list. For robust programmatic use, rely on the trajectory_url column in the statistics DataFrame, which explicitly associates each pdb_id and timepoint with the correct trajectory file.
Troubleshooting
- Statistics output too large: Error like "Statistics output too large: 75000 rows (num_steps=750000, report_interval=10). Maximum allowed: 50000 rows. Increase report_interval to at least 15." means num_steps / report_interval is too high. Raise report_interval or reduce num_steps so the ratio is ≤ 50000.
- Empty PDB input: Error "Input pdb dictionary is empty. At least one PDB structure is required." indicates that pdb was empty or unconnected. Ensure an upstream node (such as OpenMM Energy Minimize) is connected and produces at least one PDB entry.
- Simulation instability or NaNs: If backend logs mention NaNs in energies or constraint failures, lower step_size_ps, ensure suitable constraints (HBonds or AllBonds), and make sure the system was solvated and minimized upstream. For challenging systems, run shorter equilibration segments first.
- Platform or GPU errors: Errors referring to unavailable CUDA/OpenCL devices suggest the chosen platform is not supported. Change platform to "auto" or "CPU", or configure the environment with compatible GPU hardware and drivers before using "CUDA" or "OpenCL".