PDB Combiner
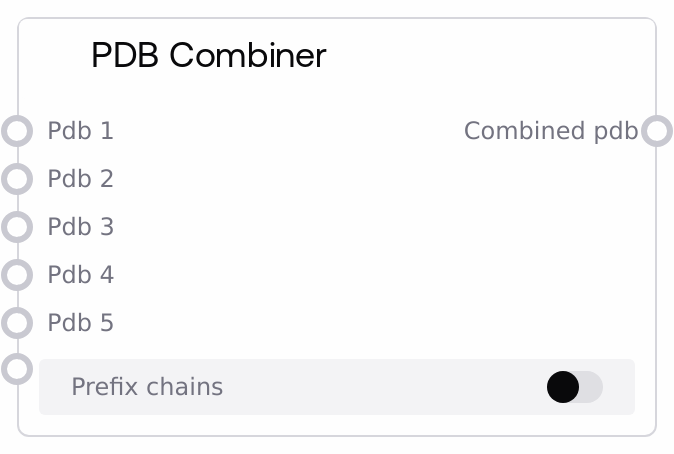
PDBCombinerNode merges multiple PDB inputs, where each is a dictionary mapping structure IDs to PDB-formatted text, into a single combined PDB dictionary. It can optionally prefix IDs per input slot (PDB1_, PDB2_, etc.) to avoid collisions when upstream nodes reuse the same identifiers. This helps assemble multi-structure datasets for downstream modeling, analysis, visualization, or export in SaltAI biotech workflows.
Usage
Use PDBCombinerNode when you have several PDB-producing steps and need a single consolidated PDB mapping for later nodes. Typical upstream producers include LoadPDBNode (user-provided structures), BatchPDBNode (batched PDB maps), PDBChainExtractorNode (subsetted chains), and PdbFixerNode (sequence-fixed PDBs). Place PDBCombinerNode after these nodes and connect any subset of pdb_1–pdb_5. Then connect combined_pdb to nodes such as PDBVisualizationNode (for 3D viewing), PDBToFastaNode (sequence extraction), PDBChainExtractorNode (further filtering), PdbFixerNode (chained fixing), SaveToZipNode, or SaveToBucketNode (export). If different inputs might share the same IDs, enable prefix_chains so each source’s IDs are namespaced (PDB1_, PDB2_, etc.). If you already enforce globally unique IDs upstream, you can keep prefix_chains false, but understand that any unintended duplicates will be auto-renamed with numeric suffixes, which changes the exact keys seen by downstream nodes. In complex workflows, you can chain multiple PDBCombinerNode instances to aggregate more than five sources or to structure your datasets by logical groups (e.g., templates, designs, controls).
| Field | Required | Type | Description | Example |
|---|
| pdb_1 | False | PDB | First optional PDB dictionary to merge. Must be a mapping from structure IDs to PDB-formatted strings, such as {"pdb_id": "PDB text"}. Typically produced by nodes like LoadPDBNode, BatchPDBNode, PDBChainExtractorNode, or PdbFixerNode. All entries from this dictionary are copied into the combined output if provided. | Example: {"1ABC_A": "HEADER TEST STRUCTURE\nATOM 1 N MET A 1 ...", "1ABC_B": "HEADER TEST STRUCTURE CHAIN B\nATOM 1 N GLY B 1 ..."} |
| pdb_2 | False | PDB | Second optional PDB dictionary to include. Same required structure as pdb_1. If keys in pdb_2 duplicate keys already seen from other inputs, they may be prefixed or renamed depending on prefix_chains. | Example: {"2XYZ_A": "HEADER DESIGNED PROTEIN\nATOM 1 N ALA A 1 ...", "2XYZ_B": "HEADER DESIGNED PROTEIN CHAIN B\nATOM 1 N LYS B 1 ..."} |
| pdb_3 | False | PDB | Third optional PDB dictionary. Often used for adding additional models, design rounds, or templates to the combined structure set. | Example: {"design_round3": "HEADER ROUND 3 MODEL\nATOM 1 N SER A 1 ..."} |
| pdb_4 | False | PDB | Fourth optional PDB dictionary to merge. Inputs do not have to be contiguous; you may leave some pdb_i unconnected. | Example: {"scaffold_A": "HEADER SCAFFOLD A\nATOM 1 N THR A 1 ..."} |
| pdb_5 | False | PDB | Fifth optional PDB dictionary, allowing up to five separate PDB sources to be merged in a single node. | Example: {"template_1": "HEADER TEMPLATE STRUCTURE\nATOM 1 N GLU A 1 ..."} |
| prefix_chains | False | BOOLEAN | If true, each ID from input pdb_i is prefixed with "PDBi_" (for example, an ID "1ABC_A" from pdb_2 becomes "PDB2_1ABC_A"), guaranteeing that IDs from different inputs cannot collide. If false, original IDs are preserved, and if the same ID appears more than once across inputs, later duplicates are automatically renamed by appending numeric suffixes (e.g. "modelA_1", "modelA_2"). | True |
Outputs
| Field | Type | Description | Example |
|---|
| combined_pdb | PDB | A single merged PDB dictionary containing all entries from the provided pdb_1–pdb_5 inputs. Keys are final structure IDs (original, prefixed with PDBi_, or suffixed with numbers to avoid collisions), and values are full PDB-formatted text strings. This unified mapping is designed to be consumed directly by downstream PDB-aware biotech nodes and export nodes. | Example: {"PDB1_1ABC_A": "HEADER TEST STRUCTURE\nATOM 1 N MET A 1 ...", "PDB2_2XYZ_A": "HEADER DESIGNED PROTEIN\nATOM 1 N ALA A 1 ...", "PDB3_design_round3": "HEADER ROUND 3 MODEL\nATOM 1 N SER A 1 ..."} |
Important Notes
- Performance: The node iterates over up to five dictionaries and constructs a new dictionary in memory. This is efficient for typical structure counts, but total memory use and processing time scale with the number and size of PDB strings being combined.
- Limitations: Only five separate PDB inputs (pdb_1–pdb_5) are supported. To merge more than five sources, either pre-merge upstream or chain multiple PDBCombinerNode instances (e.g., combine five at a time, then combine those results).
- Behavior: If all PDB inputs are missing or empty, the node raises an error indicating that at least one PDB input is required. This prevents silent creation of an empty combined dataset.
- Behavior: When prefix_chains is false and keys overlap, the node resolves collisions by renaming later entries with numeric suffixes. Downstream logic that depends on exact IDs must account for this or use prefix_chains to keep IDs predictable.
Troubleshooting
- Error about missing or empty PDB input: This occurs when none of pdb_1–pdb_5 provide a non-empty dictionary. Connect at least one PDB-typed output (for example from LoadPDBNode or BatchPDBNode) and verify that those upstream nodes are configured correctly and actually produce content.
- Unexpected IDs with numeric suffixes (e.g. '1ABC_A_1'): This means multiple inputs used the same ID while prefix_chains was false. Enable prefix_chains to namespace IDs by input index or change upstream pdb_id values so they are unique and avoid automatic renaming.
- Downstream node cannot find a specific PDB ID: The ID may have been prefixed (PDBi_) or suffixed during combination. Inspect the keys of combined_pdb (for example, by exporting with SaveToZipNode or printing in a debug step) and update downstream configuration to reference the actual final IDs.
- Type or format errors in downstream PDB consumers: Ensure you connect combined_pdb into fields that expect type PDB, not STRING or generic data. Also confirm upstream nodes produced dictionaries in the form {"id": "PDB text"}, not plain strings.