Boltz Predict¶
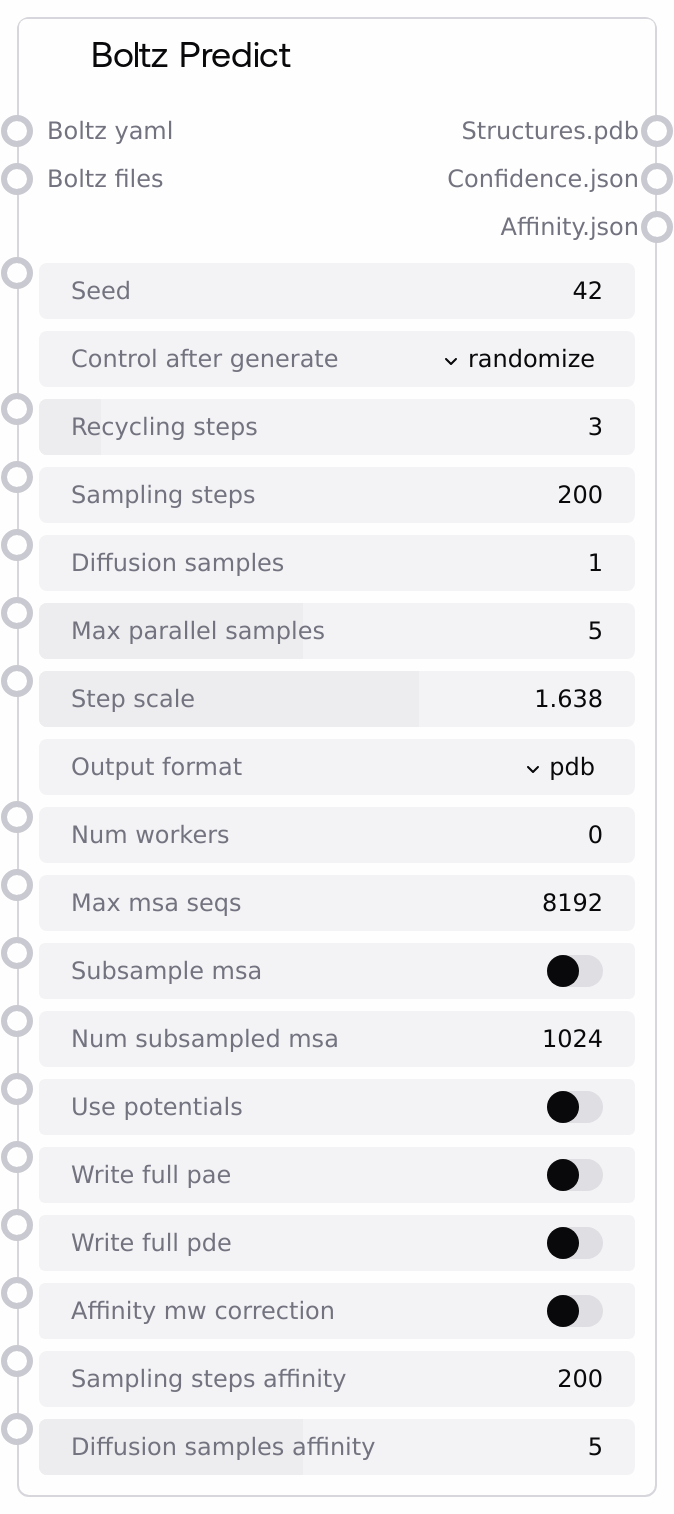
Usage¶
Use this node as the final execution step after you have constructed a valid Boltz YAML configuration and all required auxiliary files. Typical workflows start with sequence-building nodes such as BoltzProteinSequenceNode / BoltzLigandSequenceNode or the more general BoltzSequenceNode, optionally add constraints via BoltzConstraintNode, templates via BoltzTemplateNode, and properties via BoltzPropertyNode. These are combined with BoltzListCombinerNode and then passed into BoltzYAMLCombinerNode, whose boltz_yaml and boltz_files outputs should be wired directly into BoltzPredictNode.
In practice: 1) Build one or more sequences (protein, DNA/RNA, ligand) with unique chain IDs. 2) Optionally add structural constraints, templates, and affinity properties for guided design or binding affinity prediction. 3) Combine everything into a single Boltz YAML plus auxiliary MSA/template files with BoltzYAMLCombinerNode. 4) Feed both outputs into Boltz Predict and tune sampling parameters such as recycling_steps, sampling_steps, diffusion_samples, step_scale, and use_potentials.
Downstream, structures.pdb is used for visualization, analysis, or docking workflows, while confidence.json, affinity.json, pae.json, pde.json, and plddt.json feed into metrics inspection or filtering. This node talks to the Salt biotech GPU service (topic salt-saas-biotech-gpu-boltz2), so it suits non-interactive or batched structure design runs where long GPU computations are acceptable. Ensure affinity properties are only requested when at least one ligand sequence is present, and be aware that high diffusion_samples and large MSAs significantly increase runtime and resource use.
Inputs¶
| Field | Required | Type | Description | Example |
|---|---|---|---|---|
| boltz_yaml | True | BOLTZ_YAML | Boltz YAML configuration object describing sequences, constraints, templates, and properties. Must be a dictionary with at least a non-empty sequences list in the Boltz2 format, typically produced by BoltzYAMLCombinerNode. If properties includes an affinity entry, at least one ligand sequence must be present. | {"version": 1, "sequences": [{"protein": {"id": "A", "sequence": "MKTAYIAKQRQISFVKSHFSRQDILD", "_sequence_name": "protA", "msa": "msa_1.a3m", "_msa_format": "a3m"}}, {"ligand": {"id": "L", "smiles": "CC(=O)Oc1ccccc1C(=O)O"}}], "properties": [{"affinity": {"binder": "L"}}], "templates": [{"pdb": "template_1.pdb", "chain_id": "A", "template_id": "A"}]} |
| boltz_files | True | BOLTZ_FILES | Dictionary mapping auxiliary file names (MSA and template structures) to their contents. Keys are filenames referenced from boltz_yaml (for example msa_1.a3m, template_1.pdb), values are text contents for those files. Normally generated alongside boltz_yaml by BoltzYAMLCombinerNode. | {"msa_1.a3m": ">protA\nMKTAYIAKQRQISFVKSHFSRQDILD\n>homolog1\nMKTTYIAKQKQISFVKSHFSRQDILN", "template_1.pdb": "ATOM 1 N MET A 1 11.104 13.207 2.100 1.00 20.00 N ..."} |
| seed | True | INT | Random seed for the Boltz2 diffusion process. Controls reproducibility of sampling; runs with the same configuration and seed should produce identical outputs. Must be between 0 and 2^32-1. | 42 |
| recycling_steps | False | INT | Number of recycling steps in the Boltz2 model, controlling how many times the structure is refined. Higher values (for example 3–10) can improve quality but increase runtime. Must be between 1 and 20. | 3 |
| sampling_steps | False | INT | Number of diffusion sampling steps for structure generation. Higher values (for example 200–300) improve convergence at the cost of longer GPU time. Allowed range is 50 to 500. | 200 |
| diffusion_samples | False | INT | Number of independent diffusion samples (structures) to generate per run. Increasing this yields multiple candidate structures but scales runtime roughly linearly. Must be between 1 and 25. | 3 |
| max_parallel_samples | False | INT | Maximum number of diffusion samples to compute in parallel on the backend. Higher values use more GPU memory but can reduce wall-clock time. Valid range is 1–10. | 5 |
| step_scale | False | FLOAT | Diffusion step scale (akin to temperature) that adjusts step size in the diffusion process. Typical values are around 1.6; larger values (up to 2.0) increase exploration and diversity, smaller values make sampling more conservative. Range 1.0–2.0. | 1.638 |
| output_format | False | ENUM[pdb, mmcif] | File format for the generated structures. pdb returns structures with .pdb keys. mmcif rewrites those keys to .cif while keeping contents compatible with MMCIF workflows. | pdb |
| num_workers | False | INT | Number of worker processes used by the backend. Set to 0 to disable multiprocessing, or 1–8 to parallelize parts of the computation depending on backend configuration. | 0 |
| max_msa_seqs | False | INT | Maximum number of MSA sequences to consider per chain. Higher values (up to 16384) can improve signal but increase memory and runtime; lower values (for example 2048–8192) are often sufficient. Must be between 256 and 16384. | 8192 |
| subsample_msa | False | BOOLEAN | Whether to subsample MSA sequences down to num_subsampled_msa. Enable this when MSAs are very deep to control runtime and memory while preserving diversity. | false |
| num_subsampled_msa | False | INT | Target number of MSA sequences when subsample_msa is true. Must be between 256 and 4096. Typically 512–2048 provides a good trade-off between signal and efficiency. | 1024 |
| use_potentials | False | BOOLEAN | Enable inference-time potentials for improved structure quality, especially when constraints or templates are present. This increases computational cost but can yield more accurate and physically plausible models. | true |
| write_full_pae | False | BOOLEAN | If true, instructs Boltz2 to write full predicted aligned error (PAE) matrices, which are returned as JSON-decoded arrays. Useful for detailed interface and domain-boundary analysis, but increases output size. | false |
| write_full_pde | False | BOOLEAN | If true, instructs Boltz2 to write full predicted distance error (PDE) matrices, returned as JSON-decoded arrays. Recommended when you need detailed distance uncertainty information. | false |
| affinity_mw_correction | False | BOOLEAN | Apply molecular weight correction when computing affinity properties, adjusting predicted binding metrics by ligand size. Only used when affinity properties are requested in the YAML. | true |
| sampling_steps_affinity | False | INT | Number of diffusion sampling steps specifically for affinity estimation. Often similar to sampling_steps (for example 200–300). Must be between 50 and 500; used only when affinity properties are present. | 200 |
| diffusion_samples_affinity | False | INT | Number of independent diffusion samples used to estimate affinity. Higher numbers (up to 10) stabilize affinity estimates but increase cost; used only when affinity prediction is configured in boltz_yaml. | 5 |
Outputs¶
| Field | Type | Description | Example |
|---|---|---|---|
| structures.pdb | PDB | Dictionary of predicted ranked structures, mapping filenames to structure text. When output_format is pdb, keys end in .pdb (for example rank_001_model_1.pdb); when mmcif is selected, keys are rewritten to .cif. Values are full coordinate files suitable for visualization, refinement, or downstream modeling. | {"rank_001_model_1.pdb": "HEADER PROTEIN-LIGAND COMPLEX\nATOM 1 N MET A 1 11.104 13.207 2.100 1.00 78.50 N ...", "rank_002_model_1.pdb": "HEADER PROTEIN-LIGAND COMPLEX\nATOM 1 N MET A 1 10.954 13.407 2.300 1.00 76.10 N ..."} |
| confidence.json | JSON | JSON object containing confidence-related outputs from Boltz2, such as per-model scores (for example pLDDT summaries, ranking scores). The exact schema follows Boltz2 confidence output, typically keyed by model name. | {"rank_001_model_1": {"mean_plddt": 0.87, "mean_pae": 2.3, "ranking_score": 0.91}, "rank_002_model_1": {"mean_plddt": 0.82, "mean_pae": 3.1, "ranking_score": 0.85}} |
| affinity.json | JSON | JSON object with predicted affinity metrics when affinity properties are requested and a ligand is present. Empty {} when affinity is not configured. Keys usually correspond to ranked models and contain predicted binding scores or energies. | {"rank_001_model_1": {"kd_nM": 25.4, "deltaG_kcal_per_mol": -10.2, "confidence": 0.78}} |
| pae.json | JSON | Decoded PAE (predicted aligned error) matrices. The node decodes base64-encoded NPZ outputs into JSON-serializable dictionaries of arrays. The top-level keys are identifiers for each matrix; each maps to a dictionary where each array name maps to a 2D list of floats. | {"rank_001_pae": {"pae": [[0.5, 0.8, 1.2], [0.7, 0.4, 1.0], [1.1, 0.9, 0.3]]}} |
| pde.json | JSON | Decoded PDE (predicted distance error) matrices, with the same structure as pae.json. Only populated if the backend writes PDE outputs. Useful for analyzing uncertainty in inter-residue distances. | {"rank_001_pde": {"pde": [[0.3, 0.6, 1.1], [0.6, 0.2, 0.9], [1.0, 0.8, 0.4]]}} |
| plddt.json | JSON | Decoded pLDDT (per-residue confidence) arrays from NPZ into JSON-serializable form. Each key corresponds to a model, with internal keys (such as plddt) mapping to 1D lists of floats in the 0–100 range. | {"rank_001_plddt": {"plddt": [92.1, 88.4, 85.0, 90.3, 91.7]}} |
Important Notes¶
- Performance: This node calls a remote GPU-backed Boltz2 service with a timeout up to 10,800 seconds; high diffusion_samples, large MSAs, or many chains can lead to long runtimes and heavy GPU usage.
- Limitations: Affinity prediction requires at least one ligand sequence in boltz_yaml and an affinity property; the node will raise an error if affinity is requested without ligands.
- Behavior: When output_format is set to mmcif, structure keys are rewritten from .pdb to .cif but the underlying text comes from the service as ranked PDBs; plan downstream parsing accordingly.
- Behavior: PAE, PDE, and pLDDT outputs are converted from base64-encoded NPZ files into nested JSON-friendly structures (lists), which may be large for long sequences and should be handled carefully in downstream nodes.
Troubleshooting¶
- Invalid Boltz YAML: If you see errors like Boltz YAML must contain at least one sequence or Boltz YAML must be a dictionary, verify that you are connecting the boltz_yaml output from BoltzYAMLCombinerNode directly and that upstream sequence builders produced at least one valid sequence.
- Affinity without ligand: The error Affinity prediction requires at least one ligand sequence indicates that your YAML properties include an affinity block but no ligand entries in sequences; either add a ligand sequence (for example via BoltzLigandSequenceNode) or remove the affinity property.
- Type mismatch for boltz_files: If you get Boltz files must be a dictionary, check that you are wiring the correct second output from BoltzYAMLCombinerNode and not a list or a single sequence/constraint node directly.
- Timeout or empty results from backend: If the node appears to hang or returns empty structures, your job may be exceeding backend limits or failing remotely. Try reducing diffusion_samples, lowering sampling_steps, trimming MSAs (max_msa_seqs or subsample_msa), or simplifying the system (fewer chains, less complex constraints) and re-run.
- Large JSON outputs: When handling pae.json, pde.json, or plddt.json, memory issues may arise for very large systems. Consider turning off write_full_pae/write_full_pde or post-processing these outputs in specialized nodes to avoid copying large matrices unnecessarily.