Alphafold¶
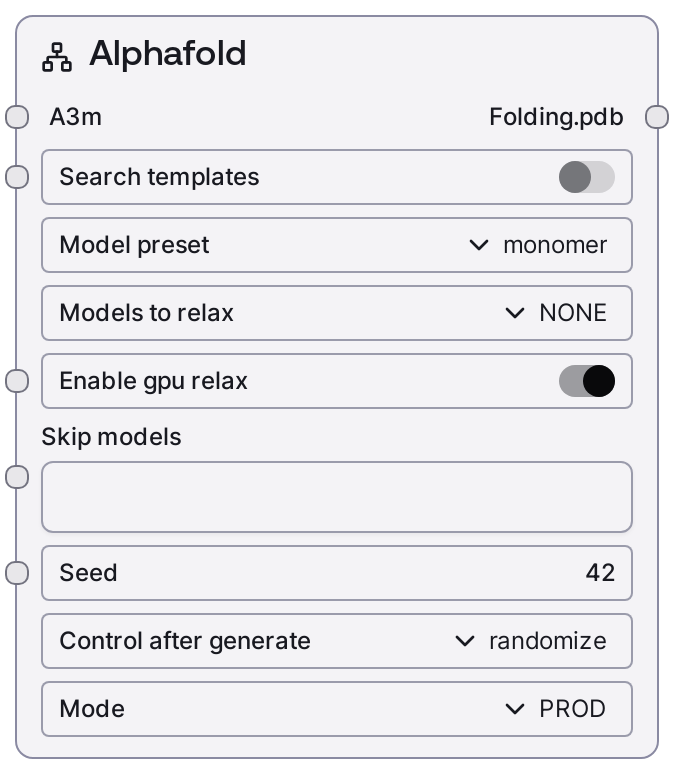
Usage¶
Use this node after generating MSAs in A3M format for your protein sequences, typically with SaaSMSASearchNode. Connect the msa.a3m output into the a3m input of SaaSAlphafoldNode, then configure model_preset, search_templates, skip_models, seed, and mode. Run in PROD for full AlphaFold predictions, MOCK for fast UI/debugging with precomputed data, or TEST for quick validation on short sequences with reduced settings.
Typical position in a workflow: 1) Upstream: FASTA creation/import → SaaSMSASearchNode (MSA Search) → SaaSAlphafoldNode. 2) Downstream: PDB consumer nodes such as structure viewers, structure-quality scoring, pLDDT/PAE analysis, docking, or design-refinement nodes.
Practical guidance: - Keep a3m keys (sequence IDs) unique and stable—they become prefixes in output keys like "SEQ1_rank_1". - Choose model_preset according to your scenario: "monomer" or "monomer_ptm" for most single-chain proteins; "monomer_casp14" for CASP-like benchmarks. "multimer" is currently rejected. - Use skip_models to skip particular model indices and reduce runtime; leave it empty to run all 1–5. - Fix the seed to obtain reproducible predictions; for multi-sequence batches, the node offsets the seed per sequence (seed + index). - Use TEST mode only for quick sanity checks on short test sequences, not as a substitute for full production runs.
Inputs¶
| Field | Required | Type | Description | Example |
|---|---|---|---|---|
| a3m | True | MSA | Multiple sequence alignments in A3M format, provided as a mapping {sequence_id: a3m_content}. Each value must be a valid A3M alignment string for one target sequence, including the query and homologous sequences in standard A3M syntax. Keys (sequence_id) must be unique and are used to label outputs; very long sequences and deep MSAs increase runtime. | {"SEQ1": ">SEQ1/1-120\nMKTAYIAKQRQISFVKSHFSRQLEERLGLIEVQAN...*\n>HOMOLOG1/1-120\nMKTAYIARQRQISFVKSHFTRQLEERLGLIEVQAN...*\n","SEQ2": ">SEQ2/1-95\nGHHHHHHSSGLVPRGSHMASMTGGQQMGRGSEF-...*\n>HOMOLOG2/1-95\nGHHHHHHSSGLVPRGSHMASMTGGQQMGRGSEF-...*\n"} |
| search_templates | True | BOOLEAN | Whether to search for structural templates before prediction. If true, the backend attempts to find homologous structures and include template features, which can improve accuracy for proteins with known structural relatives but adds overhead. If false, AlphaFold runs without template search. In TEST mode this is internally forced to false for speed. | false |
| model_preset | True | STRING | AlphaFold model preset to use. Allowed values: "monomer", "monomer_ptm", "monomer_casp14", "multimer". "monomer" is the default for typical single-chain predictions; "monomer_ptm" provides additional predicted TM-score-related metrics; "monomer_casp14" matches the CASP14 configuration. "multimer" is not supported and selecting it will raise an error. | monomer_ptm |
| models_to_relax | True | STRING | Controls post-prediction structure relaxation. Accepted values: "NONE", "BEST", "ALL". Currently only "NONE" is supported; using "BEST" or "ALL" will raise a NotImplementedError. Keep this at "NONE" unless relaxation support is added in future updates. | NONE |
| enable_gpu_relax | True | BOOLEAN | Indicates whether relaxation (if enabled in the future) should run on GPU (true) or CPU (false). Because relaxation is not currently supported, this flag has no practical effect but must still be provided; leaving it at the default true is recommended. | true |
| skip_models | True | STRING | Comma-separated list of model indices (1–5) to skip during prediction, such as "3,5". The node parses this string into integers, verifies that each index is in the range 1–5, and checks that not all models are skipped. An empty string means no models are skipped (all models run). In TEST mode, this value is overridden internally so that models 2–5 are skipped and only model 1 is used. | 2,4 |
| seed | True | INT | Base random seed for AlphaFold runs, as a 32-bit unsigned integer between 0 and 4294967295. For multiple sequences in a3m, the node uses seed + i for the i-th sequence, giving each sequence a unique but deterministic seed when re-running with the same configuration. | 42 |
| mode | True | STRING | Execution mode. "PROD" runs the real AlphaFold pipeline on managed services. "MOCK" returns predefined mock results from bundled JSON files without performing compute, useful for UI testing or development. "TEST" uses minimal parameters for speed (e.g., only model 1, no template search) and is intended for quick checks on short sequences rather than high-quality predictions. | PROD |
Outputs¶
| Field | Type | Description | Example |
|---|---|---|---|
| folding.pdb | PDB | Predicted protein structures as a dictionary mapping " | {"SEQ1_rank_1": "REMARK 1 PREDICTED BY ALPHAFOLD\nATOM 1 N MET A 1 11.104 13.207 10.423 1.00 60.00 N \nATOM 2 CA MET A 1 12.512 13.530 10.102 1.00 60.00 C \n...\nEND\n","SEQ1_rank_2": "REMARK 1 PREDICTED BY ALPHAFOLD\nATOM 1 N MET A 1 10.954 13.005 10.612 1.00 58.50 N \n...\nEND\n","SEQ2_rank_1": "REMARK 1 PREDICTED BY ALPHAFOLD\nATOM 1 N GLY A 1 5.321 8.745 3.219 1.00 62.30 N \n...\nEND\n"} |
Important Notes¶
- Performance: Runtime increases with sequence length, MSA depth, and number of sequences. The node relies on CPU and GPU services with fixed timeouts (hundreds of seconds for feature combination and up to roughly 2000 seconds for folding). For large proteins or big batches, consider splitting runs, lowering MSA depth, or using skip_models to reduce the number of models.
- Limitations: Setting models_to_relax to anything other than "NONE" or choosing model_preset="multimer" will immediately raise a NotImplementedError. The current implementation supports only monomer-style predictions without relaxation or multimer models.
- Behavior: skip_models is strictly validated. Any index outside 1–5, malformed values, or a configuration that effectively skips all models triggers an error before the request is sent to backend services. In TEST mode, the node overrides skip_models to skip models 2–5 and disables template search for speed.
- Behavior: The node converts A3M MSAs to Stockholm format and reconstructs FASTA sequences internally. If the IDs derived from FASTA and the feature IDs from the backend do not match, it raises a ValueError, usually indicating malformed A3M input or inconsistencies introduced upstream.
Troubleshooting¶
- Invalid skip_models value: If you see an error like "Expected models indices in skip_models to be in range [1, 5] but got [...]", check that skip_models is a comma-separated list such as "2,4" containing only integers 1–5 with no extra characters. Avoid using 0, 6, or text labels.
- All models skipped error: The error "Cannot skip all 5 models. Remove at least one model from skip_models" means your list covers all indices 1,2,3,4,5. Remove at least one index or leave skip_models empty so that at least one model actually runs.
- Unsupported relaxation or multimer: Errors like "Relaxation is not supported yet" or "Multimer model is not supported yet" indicate that models_to_relax must be "NONE" and model_preset must be one of "monomer", "monomer_ptm", or "monomer_casp14". Adjust these settings before re-running.
- ID mismatch or malformed input: An error about FASTA IDs not matching feature IDs suggests your A3M inputs are malformed or have inconsistent identifiers. Ensure that your a3m dict keys are simple, consistent sequence IDs and that you pass the unmodified msa.a3m output from SaaSMSASearchNode directly into this node.
- Slow or timed-out runs: If runs appear to hang or exceed expected time, you may be hitting backend timeouts due to very long sequences or large batches. Try processing fewer sequences at once, shortening sequences, reducing models via skip_models, or using TEST mode on truncated sequences to confirm the pipeline is configured correctly.